In vivo genetic screening platform
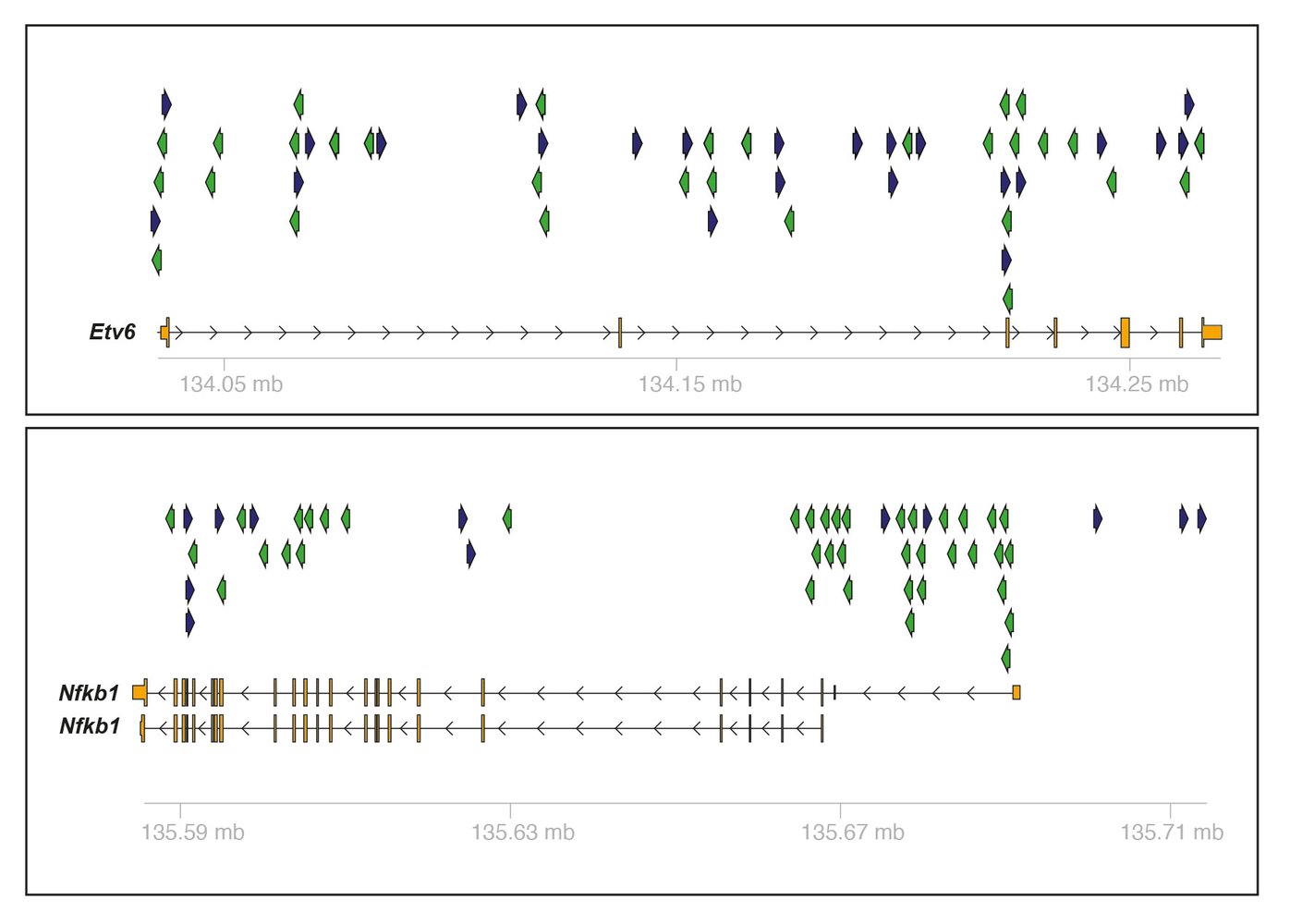
Our group has generated a series of genetically engineered mouse models, capturing a wide array of entities, ranging from indolent to highly aggressive lymphoma models. In designing these models, we aim at precisely mimicking human disease biology. In addition to serving as simple avatars of human disease, we also employ these models as backbones for in vivo genetic screens. To search for disease and drug response-modifying genes and pathways, we employ a PiggyBac in vivo insertional mutagenesis system. This screening platform delivers hits, which serve as a biological filter that helps to validate sequencing results obtained from both murine and human lymphomas and thus provides additional genetic evidence for rare variants detected by sequencing approaches.
Next to the PiggyBac screening technology, we equipped our genetically engineered mouse models with conditional Cas9 alleles to enable in vivo CRISPR screens. We deploy this technology to not only conduct lymphoma cell-specific in vivo dropout screens, but also to screen for genes and pathways that modify lymphoma immune surveillance activity and to search for modifiers of CAR-T cell fitness, expansion, exhaustion and cytotoxicity in vivo.